[ICML 2025]MindAligner: Explicit Brain Functional Alignment for Cross-Subject Visual Decoding from
计算机-人工智能-fMRI解码跨被试少样本重构

论文代码:https://github.com/Da1yuqin/MindAligner
英文是纯手打的!论文原文的summarizing and paraphrasing。可能会出现难以避免的拼写错误和语法错误,若有发现欢迎评论指正!文章偏向于笔记,谨慎食用
目录
2.3.1. fMRI-Based Brain Decoding
2.3.2. Cross-Subject Functional Alignment
2.5.3. Brain Functional Alignment Module
2.6.4. fMRI-based Visual Decoding
2.6.5. Brain Functional Alignment Analysis
1. 心得
(1)好多做跨被试的啊...
(2)这写作风格???黑人问号脸?(不是说好或者坏我只是(觉得很独特
2. 论文逐段精读
2.1. Abstract
①Limitation: model is designed for single subject
②Thus, they proposed cross-subject MindAligner
2.2. Introduction
①Limitations of alignment in cross-subject model: scarce data in shared space construction, and subject respond differently in watching the same image
②Conception of MindAligner:
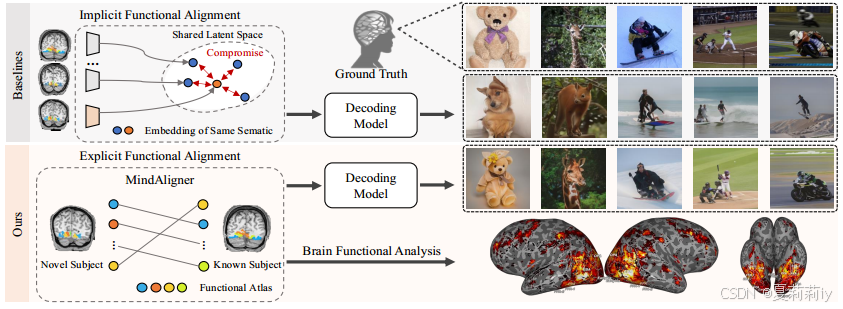
2.3. Related Work
2.3.1. fMRI-Based Brain Decoding
①Lists traditional methods and point out their work might be inefficient in cross subject reconstruction
2.3.2. Cross-Subject Functional Alignment
①Align limits inter subject specificity
2.4. Preliminary
①Only use one hour of data to pretrain, then test model in shared test set
2.5. MindAligner
2.5.1. Overview
①The overall framework of MindAligner:
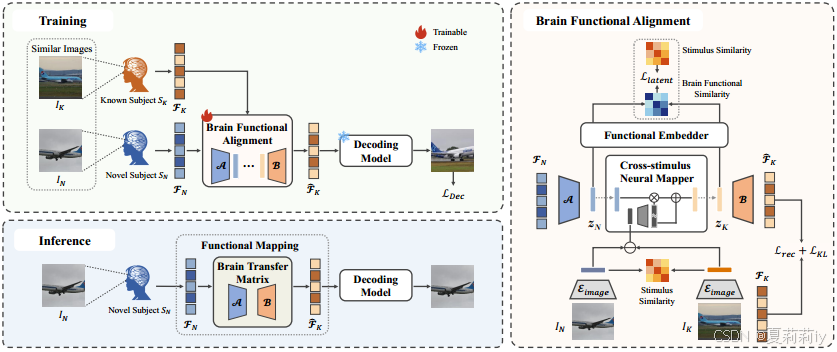
2.5.2. Brain Transfer Matrix
①Given the the fMRI signal for a subject
, the brain transfer matrix (BTM) maps it to:
②The can be decomposed to two low-rank matrices:
where and
,
and
denotes the fMRI voxel dimensions of unkown and kown subjects,
is the hidden dimension
2.5.3. Brain Functional Alignment Module
①Generate the stimuli embedding of unkown subject by stimuli differential condition to align kown embedding
:
where is pretrained CLIP, as the image encoder.
is the cross-stimulus neural mapper, 它使用
将条件
分解为缩放和移位参数??
②Further align by:
③Reconstruction loss:
and distribution loss:
④Loss between fMRI embedding pairs and stimuli pairs:
where and
are the image embeddings from CLIP,
denotes dissimilarity function
⑤The final loss:
where denotes the decoding loss in the baseline method
2.5.4. Inference
①BTM only
2.6. Experiments
2.6.1. Implementation Details
①BTM: consist of 2 linear layers with hidden dim of 4096
②Dimension of : 768
③Input and output dimension of functional embedder: 4096
④Loss coefficients:
⑤Learning rate: 1e-5
⑥Batch size: 16
2.6.2. Dataset
①Dataset: NSD
2.6.3. Metrics
①Lists metrics for performance and functional alignment measurement
2.6.4. fMRI-based Visual Decoding
①Visualization of reconstructed image:

②Quantitative performance:
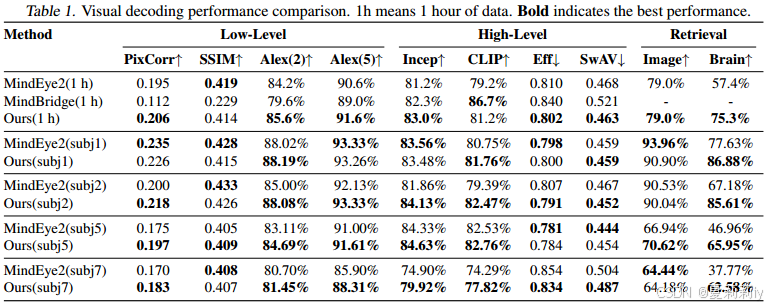
③Loss ablation:
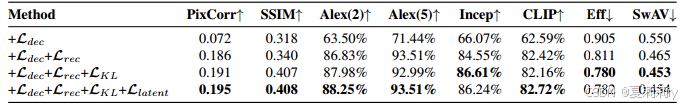
④Ablation of alignment:

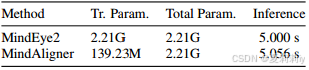
⑤Parameter comparison:
2.6.5. Brain Functional Alignment Analysis
①Visualization transfer quantity in brain:
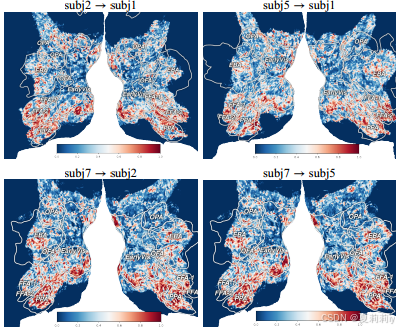
they define that early visualization region presents lower inter-subject variability, and higher visual regions (including OPA, FFA, PPA, and EBA) show larger variability
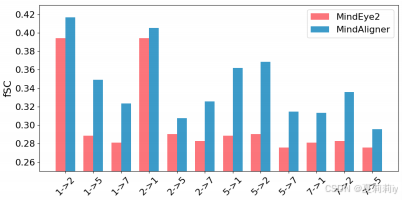
②Performance of different alignment:
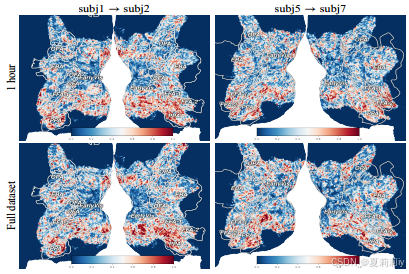
③Transfer quantity in one hour:
2.7. Conclusion
~
更多推荐
 已为社区贡献42条内容
已为社区贡献42条内容







所有评论(0)